Rapid Test Identifies Antibody Effectiveness Against COVID-19 Variants
By Ken Kingery
Test could measure patient immunity against multiple COVID-19 variants such as Omicron and Delta at once and inform which synthetic monoclonal antibody to use f

Biomedical engineers at Duke University have devised a test to quickly and easily assess how well a person’s neutralizing antibodies fight infection from multiple variants of COVID-19 such as Delta and the newly discovered Omicron variant.
This test could potentially tell doctors how protected a patient is from new variants and those currently circulating in a community or, conversely, which monoclonal antibodies to treat a COVID-19 patient. The test is described online December 3 in the journal Science Advances.
“We currently really have no rapid way of assessing variants, neither their presence in an individual, nor the ability of antibodies we possess to make a difference,” said Cameron Wolfe, associate professor of medicine at the Duke University School of Medicine. “It’s one of the lingering fears that, as we successfully vaccinate more and more people, a variant may emerge that more radically evades vaccine-induced antibody neutralization. And if that fear came true – if Omicron turned out to be a worst-case scenario – how would we know quickly enough?”
“While developing a point-of-care test for COVID-19 antibodies and biomarkers, we realized there could be some benefit to being able to detect the ability of antibodies to neutralize specific variants, so we built a test around that idea,” said Ashutosh Chilkoti, the Alan L. Kaganov Distinguished Professor and Chair of Biomedical Engineering at Duke. “It only took us a week or two to incorporate the Delta variant in our test, and it could easily be expanded to also include the Omicron variant. All we need is the spike protein of this variant, which many groups across the world — including our group at Duke— are feverishly working to produce.”
The researchers have dubbed their test the COVID-19 Variant Spike-ACE2-Competitive Antibody Neutralization assay, or CoVariant-SCAN for short. The test’s technology hinges on a polymer brush coating that acts as a sort of non-stick coating to stop anything but the desired biomarkers from attaching to the test slide when wet. The high effectiveness of this non-stick shield makes the test incredibly sensitive to even low levels of its targets. The approach allows researchers to print different molecular traps on different areas of the slide to catch multiple biomarkers at once.
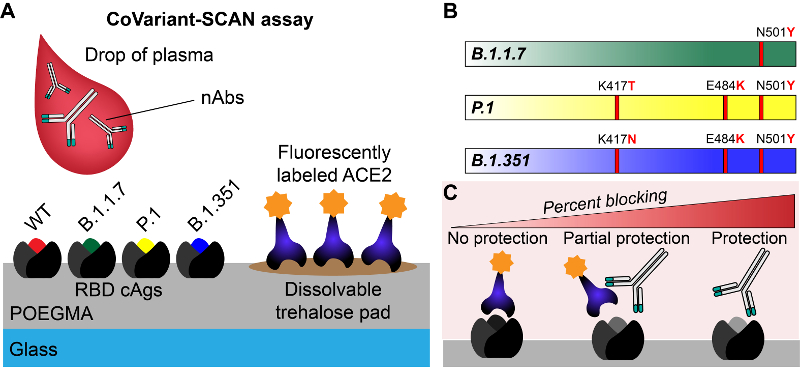 In this application, researchers print fluorescent human ACE2 proteins—the cellular targets of the virus’s infamous spike protein—on a slide. They also print spike proteins specific to each variant of COVID-19 at different specific locations. When the test is run, the ACE2 proteins detach from the slide and are caught by the spike proteins still attached to the slide, causing the slide to glow.
In this application, researchers print fluorescent human ACE2 proteins—the cellular targets of the virus’s infamous spike protein—on a slide. They also print spike proteins specific to each variant of COVID-19 at different specific locations. When the test is run, the ACE2 proteins detach from the slide and are caught by the spike proteins still attached to the slide, causing the slide to glow.
But in the presence of neutralizing antibodies, the spike proteins are no longer able to grab on to the ACE2 proteins, making the slide glow less, indicating the effectiveness of the antibodies. By printing different variants of the COVID-19 spike protein on different portions of the slide, researchers can see how effective the antibodies are at preventing each variant from latching onto their human cellular target simultaneously.
In the paper, the researchers tested the technology a number of different ways. They tried monoclonal antibodies either derived from real-life patients or from Regeneron’s commercial prophylactic treatment. They also tested plasma taken from healthy vaccinated people and those currently infected with the virus.
“In all of our tests, the results largely mimicked what we’ve been seeing in the literature,” said Jake Heggestad, a PhD student working in the Chilkoti lab. “And in this case, not finding anything new is a good sign, because it means our test is working just as well as the methods currently being used.”
While they produce similar results, the critical difference between the CoVariant-SCAN and current methods is the speed and ease with which it can produce results. Typical current approaches require isolating live virus and culturing cells, which can take 24 hours or more and requires a wide variety of safety precautions and specially trained technicians. The CoVariant-SCAN, in contrast, does not require live virus, is easy to use in most settings and takes less than an hour—potentially just 15 minutes—to produce accurate results.
Moving forward, Heggestad and the Chilkoti lab are working to streamline the technique into a microfluidic chip that could be mass produced and report results with only a few drops of blood, plasma or other liquid sample containing antibodies. This approach has already been proven to work on a similar test that can distinguish COVID-19 from other coronaviruses.
“We would love to have real-time visibility of the emerging variants and understand who still has functional immunity,” Wolfe said. “Additionally, this hints that there might be a technique whereby you could quickly assess which synthetic monoclonal antibody might be best to administer to a patient with a particular emergent variant. Currently we really have no real-time way of knowing that, so we rely on epidemiological data that can track weeks behind.”
“The reverse is also true,” Wolfe continued. “To be able to pre-screen an individual’s antibodies and predict whether they were sufficiently protected against a particular variant they are perhaps about to run into while travelling, or that is emerging in their area. We have no way of doing that at the present time. But a test like the CoVariant-SCAN could make all of these scenarios possible.”
This work was supported by the National Institutes of Health (K08HL130557, R01 AI159992, P30-CA014236, UC6AI058607), the National Science Foundation (CBET2029361), and the Department of Defense/Defense Advanced Research Projects Agency (HR0011-17-2-0069).
CITATION: “Rapid Test to Assess the Escape of SARS-Cov-2 Variants of Concern,” Jacob T. Heggestad, Rhett J. Britton, David S. Kinnamon, Simone A. Wall, Daniel Y. Joh, Angus M. Hucknall, Lyra B. Olson, Jack G. Anderson, Anna Mazur, Cameron R. Wolfe, Thomas H. Oguin III, Bruce A. Sullenger, Thomas W. Burke, Bryan D. Kraft, Gregory D. Sempowski, Christopher W. Woods, Ashutosh Chilkoti. Science Advances, Dec. 3, 2021. DOI: 10.1126/sciadv.abl7682
